
Chromium Single-Cell: Maximise data and minimise costs

The Chromium Single‑Cell Platform
The Chromium platform enables scalable, high‑resolution analysis of gene expression and cellular states at the single-cell level. By allowing researchers to understand cellular heterogeneity, discover rare populations, and link gene expression to phenotype, it provides critical insight into complex biological systems.
Designed for both discovery and translational research, Chromium supports a wide range of assays, from whole transcriptome profiling to immune repertoire and multiomic analyses, empowering researchers to confidently interrogate biology at unprecedented resolution.

Which Chromium instrument is right for your lab?
The Chromium X Series offers flexible options to match throughput, automation needs, and budget — with upgrade pathways as your research scales.
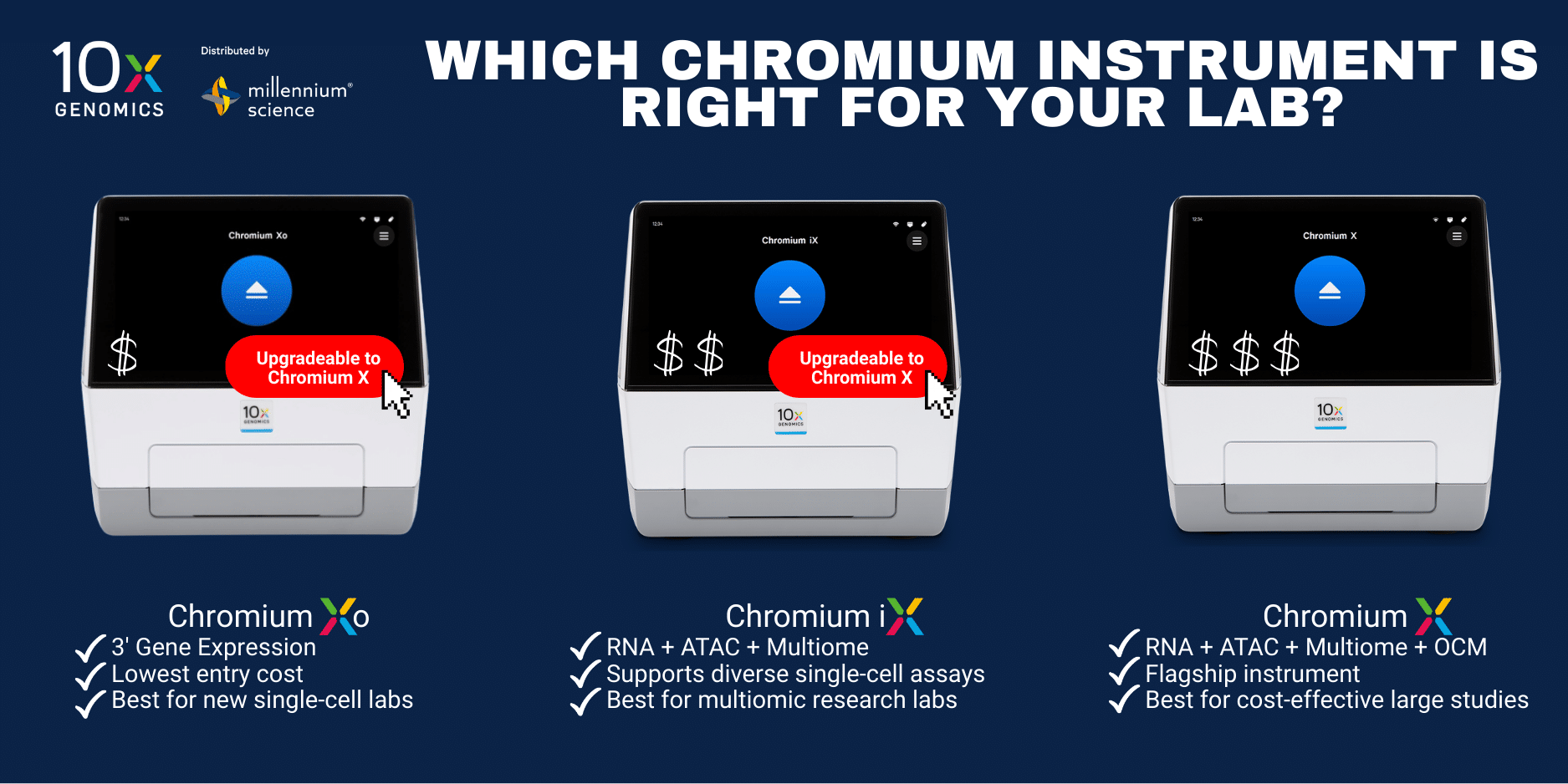

| Instrument | Assays Supported | Throughput & Workflow | Typical Use Case | Upgrade Option |
| Chromium Xo | GEM‑X Universal 3′ Gene Expression only | Moderate throughput; fast 6 min runtime | Entry‑level, efficient high‑throughput gene expression. | Can upgrade to full Chromium X |
| Chromium iX | Universal 3′ & 5′ Gene Expression, Flex Gene Expression, ATAC*, Multiome* | Moderate throughput; automated setup | Balanced throughput with expanded assay flexibility. | Upgrade to full Chromium X |
| Chromium X | Full assay suite: Gene Expression (3′/5′), Flex, ATAC, , Multiome, On-Chip Multiplexing |
High throughput; fully automated workflow | Flagship system for multiomic and high scale workflows. | N/A – flagship system |
Broad sample compatibility with integrated multiomics
Chromium assays are designed to work across diverse sample types and experimental needs, enabling seamless integration of transcriptomic, epigenetic, immune and perturbation data from the same experiment.
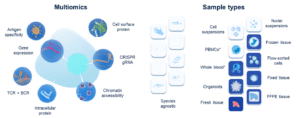
Choose the right chemistry for your samples
10x Genomics offers multiple foundational assay chemistries to accommodate fresh, frozen, fixed and FFPE samples, ensuring high-quality data even from challenging material.

Chromium Products
| Product | Core Application | Data Type | Key Features | Sample Type | Typical Use Cases |
| Chromium single-cell Gene Expression (3′ or 5′) | Single-cell transcriptomics | mRNA expression (per cell) | High-throughput, 3′ or 5′ capture, scalable | Fresh single-cell suspensions | Cell type identification, transcriptome profiling, clustering |
| Chromium single-cell Immune Profiling (5′) | Immune repertoire & gene expression | V(D)J sequences + transcriptome | Full-length TCR/BCR profiling + 5′ RNA-seq | PBMCs, lymphoid tissue | Immune diversity, clonal expansion, antibody discovery |
| Chromium single-cell Fixed RNA Profiling (Flex) | Transcriptomics from fixed cells | Targeted RNA expression | Compatible with fixed/frozen samples, up to 20k+ genes | Methanol-fixed cells or tissues | Biobanking, clinical samples, hard-to-process or archived samples |
| Chromium single-cell ATAC | Epigenetic profiling | Chromatin accessibility | Single-cell ATAC-seq, regulatory elements | Nuclei suspensions | Chromatin state analysis, enhancer usage, TF motif discovery |
| Chromium single-cell Multiome (Gene Expression + ATAC) | Multiomic profiling | RNA + chromatin accessibility | Simultaneous gene expression & epigenetic state | Nuclei suspensions | Linking transcription to regulatory elements |
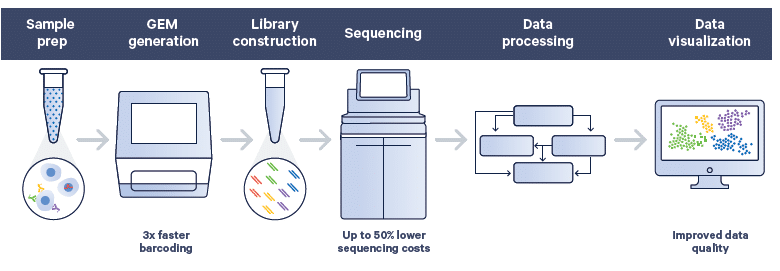
A streamlined, end-to-end workflow
Single-cell sequencing begins with a suspension of individual cells or nuclei. Using the Chromium X series instrument, advanced microfluidics generate droplets that encapsulate single cells or nuclei. Libraries are then constructed through a streamlined workflow. Once sequencing is complete, data can be processed and analysed using our integrated pipelines and visualization software, designed to be accessible for new users while providing advanced capabilities for experienced bioinformaticians.

Video of GEM (Gel Bead-In Emulsion) formation within a microfluidic channel inside the Chromium X instrument.
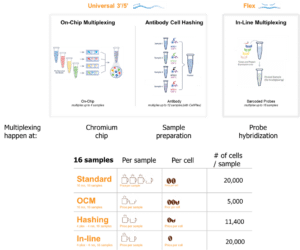
Multiplexing options
Single-cell multiplexing allows multiple samples to be analysed in a single experiment, reducing costs while increasing throughput and consistency. 10x Genomics multiplexing options help minimise batch effects and maximise the value of grant funding, making them a strong inclusion in well-designed, competitive research proposals.
| Method | Purpose | Reagents |
| On‑Chip Multiplexing (OCM) | Cost-effective sample pooling | None (chip partitioning) |
| Antibody Cell Hashing | Sample pooling upfront | Third-party Oligo‑conjugated antibodies and Cell Multiplexing Oligos |
| In-line multiplexing | Multiplex fixed samples | Probe barcodes (4 or 16) |
| Genetic Demultiplexing | In‑silico sample separation | None (Need SNPs variation) |
| CRISPR Guide Barcode | Perturbation tracking | CRISPR library w/ capture seq |
| Pre‑Index Multiplexing | Third‑party sample pooling | MULTI‑seq or custom tags |
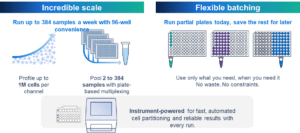
New Flex Apex delivers incredible scale
A plate-based transformation for high throughput multiplexing
– Sensitive, probe-based whole transcriptome analysis covering more than 18,000 human or mouse genes
– Fix and process samples individually (singleplex workflow) or pooled with up to 384 samples in a single lane of a Chromium chip (plate-based multiplex workflow)
– Multiomic profiling with cell surface protein, intracellular protein, or CRISPR gRNA

Single-Cell data analysis with Cell Ranger
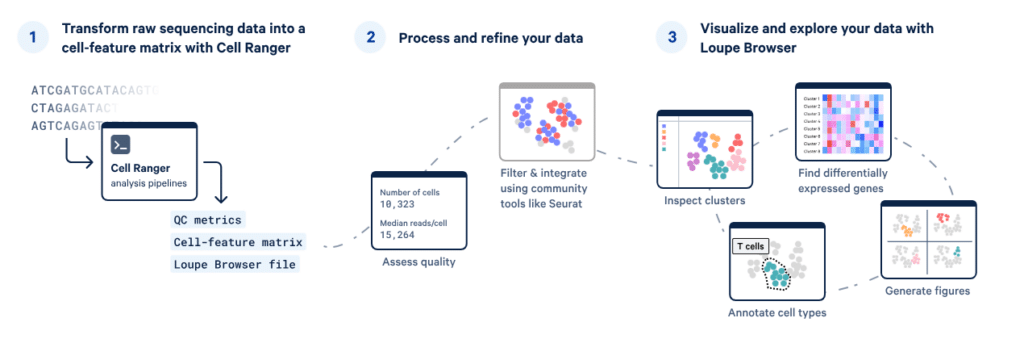
After sequencing your Chromium single-cell experiment, Cell Ranger processes raw reads to generate cell-by-gene expression matrices, perform barcode filtering, and align transcripts to a reference genome. It supports multiple pipelines, including gene expression, immune repertoire profiling, and feature barcode analysis, providing ready-to-use outputs for downstream exploration. Processed datasets can be visualized in Loupe Browser or analysed with community tools like Seurat to uncover cellular heterogeneity, cluster populations, and annotate cell types.
Cloud-Based Analysis
Cell Ranger also integrates with the 10x Genomics cloud platform, enabling secure, scalable processing of single-cell datasets without local computational infrastructure. Researchers can run standard pipelines, monitor progress, and access results from anywhere, making it easier to share datasets, collaborate across teams, and scale experiments from dozens to thousands of samples.
On this page
- Chromium Single-Cell: Maximise data and minimise costs
- The Chromium Single‑Cell Platform
- Which Chromium instrument is right for your lab?
- Broad sample compatibility with integrated multiomics
- Choose the right chemistry for your samples
- A streamlined, end-to-end workflow
- Multiplexing options
- New Flex Apex delivers incredible scale
- Single-Cell data analysis with Cell Ranger
- Request a single‑cell consultation